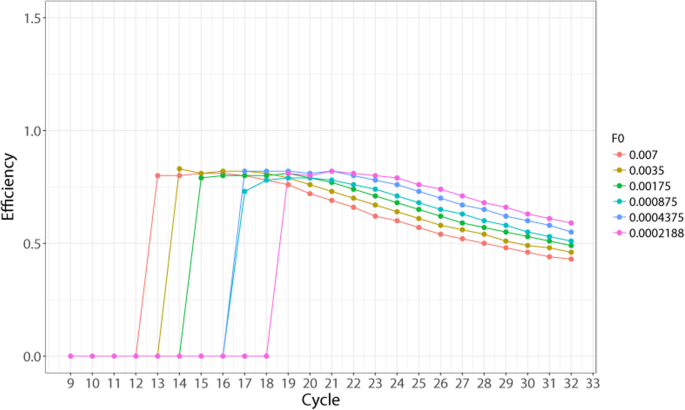
AScall processed 4,800 qPCRs in 1.5 min, with only two genotype mismatches compared to manual analysis.

Genotyping was done manually and with AScall for 96 genomic DNA samples. We show the use of AScall for SNP and indel genotyping for the MiHA study, but anyone can use the application for an accordingly designed AS-qPCR experiment of their own. With a convenient interface mimicking a regular qPCR machine interface, our tool allows batch qPCR data import via universal RDML format, amplification curves preprocessing, quality control, sample genotype calling, detailed results visualization, and report generation. Here we fill this gap and describe AScall – the interactive web application for the proposed genotyping method.
Biorad cfx manager adjust baseline software#
Earlier, we have confirmed and published the AS-qPCR method, but standard software for qPCR analysis does not suit AS-qPCR data.
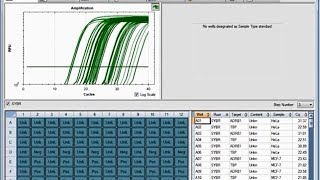
The new approach allows for multiplexing up to 480 reactions per 96 well plate and differs from common qPCR based genotyping methods. To determine the genetic mismatches between a donor and a recipient, we have implemented a real-time PCR method in conjunction with allele-specific primers (AS-qPCR). MiHAs originate from donor-recipient discrepancies in single nucleotide polymorphisms, insertions, and deletions. They may cause both beneficial and detrimental effects in forms of graft-versus-tumor and graft-versus-host accordingly. Minor histocompatibility antigens (MiHA) are critical elements for the immune response after allogeneic hematopoietic stem cell transplantation.
2Laboratory for Transplantation Immunology, National Research Center for Hematology, Moscow, Russia.1Molecular Oncology Laboratory, Pirogov Russian National Research Medical University, Moscow, Russia.These results are analyzed in more detail in “HOX genes function in Breast Cancer development” and are potentially relevant for a better understanding of the molecular heterogeneity of breast cancer, in addition to be a valuable tool assisting researchers in the choice of the most suitable cell models to perform functional assays concerning HOXB7, HOXB8 and HOXB9 genes.Konstantin Blagodatskikh 1*, Dmitrii Romaniuk 2 and Dmitry Malko 2 In addition, we found that the magnitude of the upregulation is highly subtype-specific, being the HER2+ cells the model with lowest HOXB7 upregulation, presenting very low or even null expression for HOXB8 and HOXB9, respectively. We showed that HOXB7 is upregulated in all breast cancer cells analyzed, while HOXB8 and HOXB9 are significantly upregulated in MCF7 (Luminal A), BT474 (Luminal B) and MDA231 cells (Triple-negative Claudin-low). This technique allows a very sensitive quantification of gene expression and was performed using the fluorophore SYBR Green in order to obtain the expression levels relative to a reference gene, GAPDH in this case. Here, we provided the analyses of HOXB7, HOXB8 and HOXB9 expression profiles in cell lines extensively used in the literature addressing the putative role of HOXB genes in breast cancer (MCF7, BT474, SKBR3, MDA231 and MDA468) and representative of the clinical breast cancer molecular subtypes (Luminal A, Luminal B, HER2+ and Triple-negatives Claudin-low/Basal), compared to a normal breast model (MCF10A), using quantitative-PCR (qPCR). However, due to breast cancer complexity and heterogeneity, added to the use of different technical approaches, the observed expression profiles are sometimes contradictory. Alterations in HOXB genes expression in breast cancer have been described and related to therapy response and disease progression.
